Omics Research
Transcriptomics
Transcriptomics
SARS-CoV-2 Variant Surveillance
Identify and classify all variants of concern — including the sequences representative of the Omicron and Delta — via technology designed to avoid dropouts due to mutations in primer binding regions.
What is SARS-CoV-2 Genomic Surveillance?
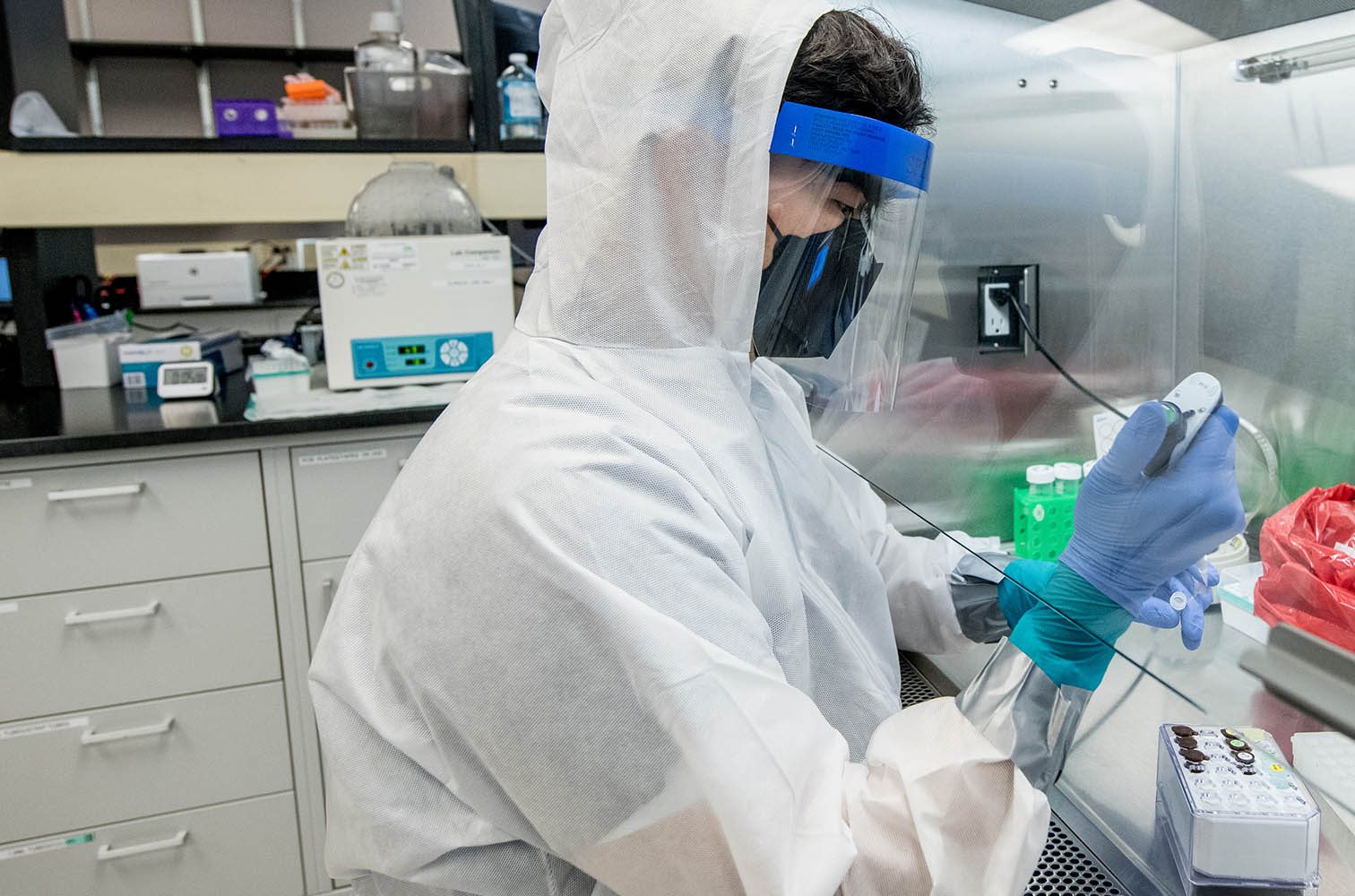
At Psomagen, SARS-CoV-2 Variant Surveillance is a genome-wide variant characterization of the virus known to cause COVID-19. As new strains of this novel coronavirus appear over time, surveillance efforts serve public health interests by tracking break-outs as well as transmission and mutation rates. This analysis includes further classification of specific variants within diverse sample types, including wastewater, and is an important defense for outbreak containment.
Why is Surveillance Important?
Use our IDT® xGen™ SARS-CoV-2 Expanded Amplicon Panel to generate a readout of all variants of concern, including Omicron and Delta. This amplicon sequencing panel contains overlapping primers and super amplicon technology that is designed to avoid dropouts (non-amplification due to mutations in primer binding regions). You can be confident that our panel will amplify not only the latest variants but those that emerge in the future.
What Can You Do?
coronavirus
Detect Novel Variants
Identify mutations via amplicon technology that avoids non-amplification (dropouts) due to mutations in primer binding regions.
gavel
Perform Government Reporting
Obtain data that can be used for public health tracking as requirements continue to evolve.
SARS-CoV-2 Variant Identification With IDT®
xGen™ SARS-CoV-2 Expanded Amplicon Panel, 96 rxn
- Best for:
Monitoring virus evolution and confidently identifying variants for strain classification to inform vaccine development, diagnostic testing, and mitigation strategies. - Benefits:
Overlapping primers provide full coverage of target regions to ensure that all variants are identified, even when those variants are deletions.
→ Learn More About the xGen™ Panel
Quotes From Our Clients
"Collaboration between vendors, like IDT and Psomagen, is critical, not only to support researchers like myself in... combating more global threats, such as the pandemic."
Arvind Varsani, Ph.D.
Arizona State University
looks_one
Request a Consultation or Quote
Get in touch with your local field team and start designing a project. We’ll help drive your research forward with the right tools for your investigative needs.
looks_two
Create, Manage and Track Your Project
At this step you'll be able to keep an eye on all the various methods and platforms we’re using to ensure your starting material will produce the most in-depth information possible. It’s easy to get in touch with us if you have a question or concern.
looks_3
Get Your Data and Analysis Reports
Securely access your raw data and view detailed analyses specific to your scientific question all in one place. Then reach out to our bioinformatics scientists to dig deeper into the results.
looks_4
Go to Our Customer Portal
Ready to get started on a new project or check in on an existing one? Do that — and more in our customer portal.
SARS-CoV-2 Diagnostic Testing
Whether you have surveillance sequencing, diagnostic, or at-home testing needs, please consider our other SARS-CoV-2 support services.
coronavirus
Psoma COVID-19 RT Test
A highly accurate SARS-CoV-2 swab test for patients traveling or returning to work or school.
Detect RNA of the SARS-CoV-2 nucleocapsid gene (N1 and N2 target) and the RNase P gene in upper respiratory and bronchoalveolar lavage specimens with this in vitro lab-developed test (LDT) that uses real-time reverse transcription-polymerase chain reaction (RT-PCR).