Defining Personalized Medical Care
Every person’s genetic structure is unique. Therefore, successful medical care is not a one-size-fits-all solution. Instead, personalized medical care shifts the focus from what works for some to what works for the individual.
DNA sequencing insights help tailor personalized medical care to the needs of the patient. The technology helps clinicians analyze the patient genome for variants, mutations, and anomalies that are indicators of disease or other health concerns.
In This Article:
- Why Personalized Medicine?
- The Role of Genetics in Disease
- DNA Sequencing and Disease Management
- Current Challenges with DNA Sequencing
Why Personalized Medicine?
Without personalized medicine, many treatment decisions are made in the dark. If a therapeutic intervention does not work for a patient, providers move on to the next available option, which can delay a patient’s receipt of proper treatment. It can also be more difficult to detect a disease in its early stages without the benefit of genetic testing.
Personalized medicine, in contrast, gives clinicians the insight they need to make targeted treatment decisions. With personalized medicine:
- Care is tailored to the patient and the genetic indicators in their genome. Gene-disease relationships can be identified in the patient’s sequencing results.
- Data from DNA sequencing can determine a person’s risk of disease before symptoms appear.
- DNA sequencing can provide care insights before, during, and after patient diagnosis.
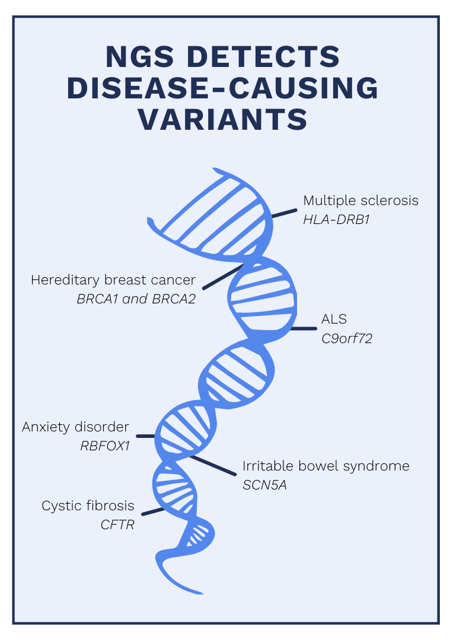
The Role of Genetics in Disease
Diseases like cancer and diabetes are associated with genetic variations in the human genome. Although two people may have the same diagnosis, the presence, manifestation, or treatment efficacy for a specific disease may vary on a genetic level.
Our knowledge of genetic links to disease continues to grow. Genetic database DisGeNET, for example, records genetic indicators of over 30,000 diseases, and genetic variants associated with over 14,000 diseases. These databases are constantly evolving, with more links being added all the time.
DNA Sequencing and Disease Management
Disease prevention and risk assessment
The role of DNA in medicine has grown exponentially in recent years. We know more about genetic predictors of disease than ever before. This makes personalized medicine more accessible for preventative medicine.
DNA sequencing allows clinicians to better understand the potential for disease, as well as prevention and treatment options based on the individual patient’s data. This also includes the benefit of becoming familiar with numerous genetic variants in different diseases.
Utilizing the markers in a patient’s genome also aids in determining what hereditary diseases they may or not be susceptible to as a means of risk assessment.
By completing DNA sequencing and analyzing the results, clinicians can:
- Target genetic variations, even for rare and/or hereditary disease, before it manifests.
- Prevent the manifestation of a disease with early intervention.
- Focus on preventive medicine options, treating a disease before it manifests or reaches more advanced stages.
- Prepare the patient for what will happen if or when the disease manifests.
- Improve the quality of life for their patients by reducing health care costs and providing a higher chance of survival.
An example of using DNA sequencing for disease prevention and management is found in a study for Spinal Muscular Atrophy that utilized gene therapy options for the disease. For some background, Spinal Muscular Atrophy degrades muscle tissue and causes neurogenic conditions in patients. For children who are diagnosed with the disease, it is often fatal. Historically, treatment options involved management of symptoms, not for the disease itself.
With DNA Sequencing technology, Spinal Muscular Atrophy has been attributed to a deficiency of the SMN protein found within the genome. Though, it is also through DNA Sequencing that researchers have formed a drug called nusinerson that alters the malfunctioning SMN protein.
Nusinerson is not the only promising treatment for those afflicted with Spinal Muscular Atrophy either. DNA Sequencing has also allowed an emergence of a new gene therapy called AVXS-101. In a separate study, AVXS-101 “increased the probability of survival, rapidly improved motor function, and enabled motor milestone achievement in SMA1 infants.”
Since the publication of both studies, AVXS-101 and nusinerson have been FDA approved to treat Spinal Muscular Atrophy.
DNA sequencing provides risk determination of disease on an individual level. In a pilot study of whole-genome sequencing in clinical applications, one in five “healthy” adult participants had a genetic alteration suspected to cause disease. Every participant had a recessive mutation associated with disease.
This provides a better picture of patient risk than traditional assessment methods. When participants only submitted their family health history, 16% of them were referred for genetic counseling or additional testing. Once participants’ genomes were sequenced, that number jumped to 34%.
Those who have higher risk of a disease may also undergo enhanced screening. After DNA analysis, for example, a patient can undergo chemoprevention to reduce their risk of breast cancer if they have a higher risk level.
Early detection
Next-generation sequencing technologies, like circulating tumor DNA analysis (ctDNA), can detect indicators of disease years before people typically receive a diagnosis. In cancer treatment, this time frame is crucial to patient outcomes. Faster diagnostics contribute to peace of mind for the patient and a timely plan of action.
The speed of early detection is a major decision-making factor for clinicians considering NGS. An example of the equipment, HiSeq X, produces up to 1.8 terabases of data in three days. This equipment, combined with a bioinformatics program, is a powerful tool for obtaining massive amounts of useful information in personalized medicine.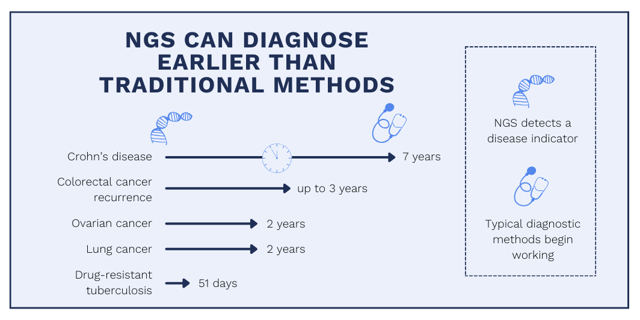
Personalized diagnostics
DNA sequencing is used regularly in diagnostics, especially in oncology. It is especially valuable in characterizing tumors. This helps clinicians select a treatment that will terminate tumor growth with the least amount of stress to the patient.
For example, genetic researchers at Washington University had a colleague with acute lymphoblastic leukemia. His tumor had not responded to chemotherapy or a bone marrow transplant. So, they sequenced the DNA of his cancer cells and healthy cells, as well as his RNA.
Sequencing data uncovered overexpression of the FLT3 gene. Treatment with sunitinib (an FLT3 inhibitor) led to success, even though the drug had only been used in kidney cancer previously. The patient achieved remission, when all typical treatments for the disease had failed.
Tailored therapeutics
Therapeutic targeting significantly reduces suffering and loss of time spent on ineffective treatments. By using therapeutic targeting, patients also avoid adverse reactions to other treatments.
DNA sequencing for therapeutic targeting can:
- Reveal information regarding genetic variations that affect drug metabolism.
- Reveal the underlying mechanism of a disease so that the mechanism can be initiated early while the patient is being treated.
- Improve the quality of life of patients.
For example, diagnosis of male breast cancer is challenging due to lack of screening and education. A male breast cancer study found that nearly 80% of men did not know men could get breast cancer. However, the link to breast cancer in men is largely due to the BRCA2 mutation found within the genome.
With specific genetic mutations contributing to male breast cancer and the lack of education that put men at risk, the study recommends targeted therapeutics for treatment partnered with conventional therapy for men’s mental wellness while undergoing treatment.
In cancer treatment, tailored therapies can include:
- Chemotherapy
- Surgery
- Immunotherapy
- Hormonal therapy
- Radiation therapy
- Molecular therapy
- Prescription medications
In a study on lung cancer, Washington University researchers performed NGS assays of genes known to be mutated in different forms of lung cancer. It took less than a month to obtain the genomic data, which revealed that 46% of the samples tested had KRAS mutations. With this information, it was possible to begin personalized therapeutic interventions. 
Choosing optimal therapies
Understanding specific DNA sequences and the genetic information of a patient can improve the odds of successful treatment by choosing optimal therapies. As discussed previously, tailored therapies include chemotherapy, surgery, immunotherapy, hormonal therapy, radiation therapy, molecular therapy, and prescription medications.
Optimal therapy includes tailored therapies by deciding what the specific components are to create, control, and regulate a regimen for specified criteria between a patient and clinician to achieve positive outcomes before, during, and after disease manifestation. How the optimal therapy is chosen is largely based on the clinician using methods to ascertain what will work best for the patient as well as clear communication with the patient on the treatment options that will most benefit them.
Current Challenges with DNA Sequencing
DNA sequencing presents some challenges within clinical medicine and management. The use of DNA sequencing requires insurance policy approval, because genetic tests are sometimes considered medical devices. If an insurance policy does not cover genetic sequencing, it is an out of pocket expense.
Pharmaceuticals designed based on DNA sequencing data must go through trials and approvals, which can take some time. So even if a genetic risk factor is identified, there may not be an approved therapeutic intervention at this time. Clinical trials are still one of the more common ways patients get access to targeted drugs.
These challenges limit the use of DNA sequencing for routine checks and personalized medicine. However, the more research that goes into personalized medicine applications of NGS, the more accessible the benefits will be to patients. DNA sequencing will enhance diagnostic and treatment efforts.
With DNA sequencing, clinicians can and will:
- Minimize misdiagnosis
- Detect disease earlier
- Use tailored therapeutics more efficiently, with lower stress and toxicity to the patient
- Improve survival rates for patients
Interested in genomics in personalized medicine? Take a look at some of our other posts:
Whole Exome Sequencing: 12 Must-Ask Questions for Clinicians >